Required Settings
Parameter Settings: Bounds
An upper and a lower limit can be defined for the parameters. A good approach is to use a very low and very high value initially that does not impose a restriction to the optimization, this is often useful in classic calibration. Alternatively, the bounds can be set according to the expected reasonable parameter ranges, PEST / PEST++ offers.
Optimization Control: Ensemble Smoothing
Applies PEST-IES to create an ensemble of a specified size, by randomizing the parameter values according to the specified prior uncertainty range and calibrating each ensemble member. The resulting ensemble yields non-linear uncertainty distributions of the parameters and can be deployed for calculating forecast ensemble runs that covers the uncertainty of the predictions. PEST-IES is also more tolerant to model instabilities and often requires much less computational time then the classic Estimation mode.
Necessary Settings
Number of Realizations (ies_num_reals)
The number of realizations defines the size of the ensemble (the number of different models that the ensemble contains). This number should exceed dimensionality of the solution space (see PEST++ Handbook, chapter 9.1.2).
Enforce Bounds
If this option is set, PEST will truncate the (log) normal distribution at these bounds. This is generally recommended as extreme parameter values may affect the stability of the underlying FEFLOW model, leading to long run-times or model failures.
Uncertainties
This defines the prior uncertainty distribution of the parameter values. The parameter values are assumed to have a normal (gaussian) distribution if no transformation is set, or log-normal distribution if log-transformation is set.
There are three principal options for setting the spread of the (log)-normal distribution:
-
bounds: The standard deviation is derived from the parameter bounds. This is useful if the parameter bounds have been defined already with the expected standard deviation of the (log) normal distribution in mind. The user must specify to how many standard deviations the interval between the bounds corresponds to (par_sigma_range). Typical values would be 4 standard deviations (relates to approx. 95% of all values) or 6 standard (relate to approx. 99.7 % of all values).
-
manual: The standard deviation is directly specified for each parameter. Pressing the … button will open a text editor, in which the standard deviation is provided for each parameter separately.
-
cov. matr: Use a covariance matrix for the generation of the ensembles.
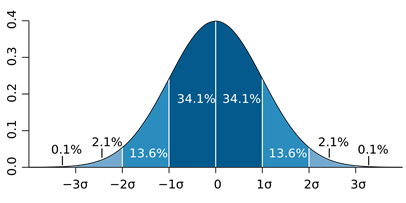
Parameter histogram.
By default, PEST++ adds a measurement noise to the observation set of each ensemble member. By default, the noise applied for each observation is has a standard deviation of 1 by the observation weight. (PEST assumes that the observation weights have been assigned using this rule when using the Ensemble Smoother with default settings).
Removing the flag will suppress the application of additional noise to the calibration dataset (ies_no_noise()).
Localisation
Because the Iterative Ensemble Smoother often explores a very large parameter space with limited amounts of models, it is unavoidable to find “spurious” (non-plausible) correlations between parameters and observations and which deteriorate the approximation of the Jacobian Matrix. Localisation informs PEST about parameter combinations that are unlikely to be correlated (e.g. distant pilot points) which improves the search progress. See the respective section in the PEST++ user manual for more information on localisation .
Three settings are available:
-
None: Do not apply localization
-
Auto: Use correlation-based, automatic adaptive localization (ies_autoadaloc()). The number of significant standard deviations (ies_autoadaloc_sigma_dist()) represents the number of standard deviations from the background mean that the value of an estimated correlation coefficient must be for it to be considered significant, as outlined in the PEST++ documentation.
-
Distance: Use localization based on distance. Correlations between observations and parameters that are separated by more than a specified distance will be considered non-plausible and ignored. The distance can be specified either as an absolute number [in meters] or relative to the average spacing.
-
Manual: Use the option for editing the localization matrix directly.